What is mtDNA?
mtDNA stands for “mitochondrial DNA”. All of us, both males and females, carry mtDNA. mtDNA is found in most of the cells in our body.
Most of the DNA in our body is found in the nucleus of our cells, but mtDNA is unique, as it is found in small structures or organelles called mitochondria. Mitochondria are found in the cytoplasm of our cells, NOT in the nucleus.
Multiple mtDNA copies in each of our cells
Many mtDNA copies are found in every mitochondria, and there are many mitochondria present in the cytoplasm of each cell. This means that we have many more copies of mtDNA in our cells than nuclear DNA, which is present in only one set per cell. The huge abundance of mtDNA, as well as its small size, make it an excellent candidate for forensic studies of old or degraded samples.
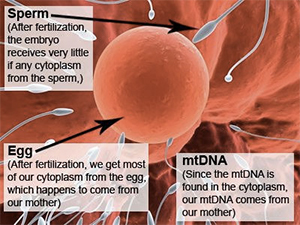
mtDNA has a unique inheritance pattern
Most of our DNA is inherited in equal proportions from each of our parents – one copy of each chromosome from each parent. But mtDNA has a unique inheritance pattern. We inherit all our mtDNA from our mother, and our mother inherited all her mtDNA from her mother, and so on. It is passed down strictly along the direct maternal line from a mother to all of her children. Males will carry the mtDNA of their mother, but when they have children, their children will carry the mtDNA of their own mother, not their father. Thus, only daughters pass the
The reason for the maternal inheritance pattern of mtDNA is due to its localization in the cell. When an egg is fertilized, the cells of the resulting embryo contain the cytoplasm of the egg, not the sperm. Since mtDNA is found only in the cytoplasm, all of our mtDNA comes from our mother, not our father. As the embryo continues to develop into a full grown human, all of the cells in the resulting human contain strictly the cytoplasm and mtDNA of the mother, not the father.
How does mtDNA hold maternal ancestral information?
The maternal inheritance pattern of the mtDNA has important significance for ancestral studies. While most of the other DNA in our body is mixed from generation to generation, mtDNA remains unmixed because it follows a strict single line of descent from mother to child. This means that our mtDNA is the same as our mother’s and our mother’s mother’s mtDNA, back thousands of generations. mtDNA testing allows us to trace our direct maternal lineage (mother’s mother’s mother’s….. maternal lineage).
Facts about mtDNA
A good understanding of the basics of mtDNA helps you to better understand mtDNA ancestry discussions in this tutorial.
What does mtDNA look like?
1. It’s round! Unlike our nuclear DNA, which is linear, mtDNA is a round circle, called a plasmid.
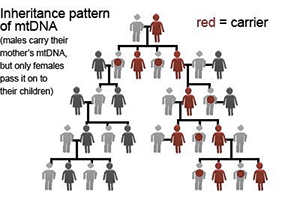
2. It’s small! While nuclear DNA is a ~247,200,000 bases pairs, mtDNA is only ~16,569 base pairs. Don’t worry if you don’t know what a base pair is – we will be talking about base pairs in more detail later in this tutorial.
Why is mtDNA so different from the other DNA in our body?
The strange appearance of mtDNA in comparison to the other DNA types in our body has something to do with its ancient origins. Mitochondria has many of the same features as single cell organisms called “prokaryotes”. Bacterial cells are prokaryotes. The mtDNA that is found inside the mitochondria is a circular plasmid, just like the DNA in bacteria.
The “endosymbiotic hypothesis” suggests that the reason for the extremely close resemblance of mtDNA to bacterial DNA is that 1.7 to 2 billion years ago, mitochondria were free-living bacteria that were “engulfed” by a cell and became permanently incorporated in the cytoplasm of the cell. This is called a “symbiotic” relationship because the cell and the bacteria provided a survival advantage to each other (mitochondria produces energy “ATP” for the cell, and the cell provides protection). This explains why the mtDNA is small and circular and found in the cytoplasm instead of the nucleus of the cell.
What does mtDNA do?
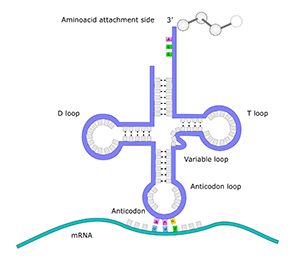
mtDNA contains the genetic code for at least 37 essential genes. 13 of the genes are responsible for producing proteins, 22 of the genes hold the genetic code to produce transfer RNA (tRNA), and two genes encode ribosomal RNA (rRNA). Thus, the mtDNA is very important, and when something goes wrong with the mtDNA, it can lead to mtDNA diseases, such as exercise intolerance or Kearns-Syre syndrome.
The size, structure and importance of mtDNA for survival, all play a role in where the majority of ancestral markers are located in the mtDNA.
mtDNA structure
mtDNA is a circular loop of DNA. DNA looks like a long ladder twisted into a double helix. The sides of the ladder are the ”backbone”, and the rungs of the ladder consist of “nucleotide bases”. There are four types of bases: T, C, A and G. T is always paired to A, and C is always paired to G. This pairing leads to the term “base pairs”.
The mtDNA loop is ~16,569 base pairs in length. The location of each base pair in the mtDNA can be specified with an accession number according to its position. When numbering the base pairs, we start at the “origin”. The origin is arbitrarily located in the D-loop between the HVR1 and HVR2 regions.
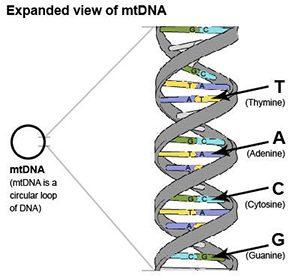
mtDNA contains three different regions
The mtDNA genome can be divided into three main regions: HVR1, HVR2 and the Coding Region. The HVR1 region is ~500 nucleotides, the HVR2 region is ~600 nucleotides, and the coding region is ~15,500 nucleotides.
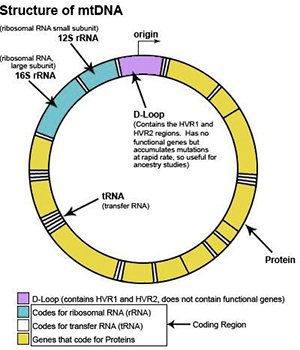
The HVR1 and HVR2 regions of the mtDNA contain the most variation, and are the most common starting point for maternal ancestral studies. The HVR1 and HVR2 regions are considered non-vital parts of the mtDNA, because they do not have a useful biological function. Thus, when a change (mutation) occurred in the HVR1 or HVR2 region of one of our ancestors, the individual did not die, and survived to pass the mutation along to all future generations.
In contrast, the coding region contains many essential genes, so when a mutation occurs in the coding region, it is often lethal. Thus, very few mutations in the coding region are passed down to future generations. For this reason, over tens of thousands of years, many mutations have accumulated in the HVR1 and HVR2 regions, but a much smaller number are found in the coding region. When tracing ancestry, scientists usually begin by testing the HVR1 and HVR2 regions because of their small sizes and abundance of mutations.
What variation occurs in the mtDNA?
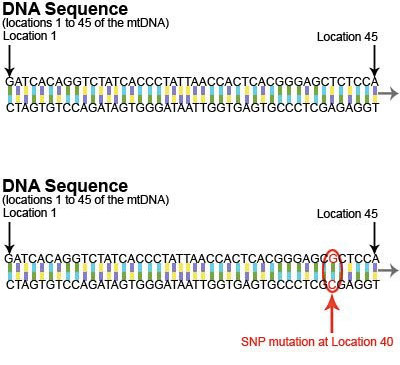
There are many types of mutations, but the type of mutation most commonly found in mtDNA is called a “SNP” (single nucleotide polymorphism). A SNP occurs when a single nucleotide is replaced with a different nucleotide. For example, in this diagram, the “T” at location 40 is replaced by a “G”.
This mutation is documented as follows:
Location: 40
Nucleotide Change: T>G (also indicated as T40G)
When you test your mtDNA, your results report will document the SNPs that you carry in your mtDNA. These are all variations from the revised Cambridge Reference Sequence.
What is the revised Cambridge Reference Sequence?
The Cambridge Reference Sequence (CRS) is a fundamental part of mtDNA data analyses.
The original CRS was the first human mtDNA genome that was fully sequenced and published. The work was performed by scientists at Cambridge University, and this groundbreaking study was officially published in 1981. Subsequent research later identified multiple discrepancies in this sequence, and in 1999 a revised Cambridge Reference Sequence (rCRS) was published.
This rCRS is now the “reference” sequence that all other mtDNA sequences are compared to. So when we state that we have variations or mutations in our mtDNA, we are actually identifying regions of our DNA which differ from the rCRS. Let’s take a look at an actual mtDNA report:

This report shows six variations in this individual’s HVR1 region, meaning that their HVR1 sequence differs from the rCRS at six different locations. For example, the 16126 T>C substitution means that the individual’s mtDNA is different from the rCRS at location 16126. It shows that the rCRS has a “T” at this location, but the person tested has a “C”.
The key point to remember is that when the results of mtDNA testing are used for genealogical purposes, the results are compared to the rCRS and mutations are reported as “differences” between the results and the rCRS. However, this can lead to confusion for beginner genetic genealogists, because instinctively people often think that when scientists look for mutations, they should be comparing our mtDNA to that of the earliest human DNA to see how our DNA has changed over time. However, that is not how the research community has decided to approach the mtDNA. The consensus within the scientific community was that mtDNA would always be compared to the rCRS.
What if I don’t have any mtDNA SNPs in my report?
If you do not have any SNPs showing in your report, that means that your mtDNA sequence (at least the part that was tested) is exactly the same as the rCRS. The rCRS belongs to a branch of haplogroup H, so if you belong to haplogroup H, chances are that you will not have many mutations in comparison with the rCRS.
How are mtDNA SNPs detected?
The genetic sequence of the mtDNA is determined using a method called “Sanger Sequencing”. This technology allows the lab to “read” the genetic code of a specified section of your mtDNA. The benefit of Sanger Sequencing is that it can accurately read entire lengths of your mtDNA. It represents the most comprehensive way to test mtDNA, as any and all SNPs will be detected. The drawback of Sanger Sequencing is that only approximately 400 to 800 nucleotides can be read at a time (in one run) so it is very expensive to perform.
Tracing ancestry with mtDNA
We all have a unique pattern of SNPs in our mtDNA, known as our mtDNA profile. This profile can be used to trace maternal ancestry in two ways – by direct comparisons and by mtDNA haplogroup and subclade determination.
Direct mtDNA profile comparisons can be made between two or more individuals to determine if they are from the same maternal lineage. Alternatively, you can use your mtDNA profile to search the DNA Reunion database to find other individuals from around the world that are from the same maternal lineage.
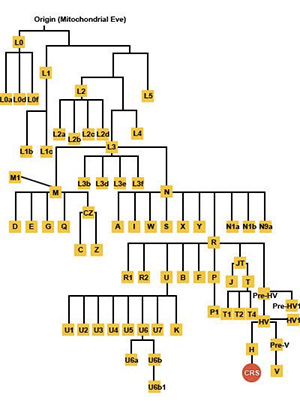
What are mtDNA haplogroups and subclades?
mtDNA studies have shown all people living today can be traced back to a common maternal ancestor (Mitochondrial Eve) who lived in Africa approximately 150,000 years ago. Over time, man journeyed out of Africa and populated the rest of the world. As these migrations took place, SNPs occurred in the mtDNA, and were passed on to all the descendants of that individual through the maternal line. These “family groups” are known as mtDNA haplogroups, and are the major branches of the mitochondrial “family tree”. The finer branches of this tree, are known as mtDNA subclades.
mtDNA haplogroups are not country specific, but specific haplogroups are more common in certain regions of the world. Hence, once you have determined your mtDNA haplogroup, you can determine which region of the world your maternal ancestors originated from.
This chart shows the mtDNA haplogroups found in each region.
| Region/Population | Major mtDNA haplogroups |
| Native Americans | A, B, C, D, X |
| Oceanic and Aboriginal Australians | P, Q, R, |
| East Asian | A, B, C, D, E, F, G, M, Y, Z |
| South Asian (i.e. India) | G, M, R, W |
| Europe and Middle East | H, HV, HV0, I, J, JT, K, R0, T, U, V, W, X |
| African | L0, L1, L2, L3, L4, L5, L6 |
mtDNA tests
There are three main mtDNA tests:
1. HVR1 Test (sequencing the HVR1 region)
2. HVR1 and HVR2 Test (sequencing the HVR1 and HVR2 regions)
3. HVR1, HVR2 and Coding Region Test, aka “mtDNA full sequencing test” (sequencing the entire mtDNA genome, which includes the HVR1, HVR2 and coding regions)
As more regions of the mtDNA genome are sequenced, the strength of the search results increases. Furthermore, sequencing of the entire mtDNA genome is required to confirm both your mtDNA haplogroup, as well as your mtDNA subclade.